Research Archive
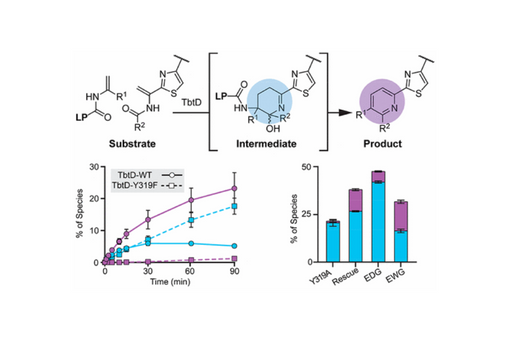
Building on our recent observation of the Bycroft–Gowland intermediate (i.e., the direct product of the [4 + 2]-cycloaddition), we interrogated thiopeptide pyridine synthases using a combination of targeted mutagenesis, kinetic assays, substrate analogs, enzyme–substrate cross-linking, and chemical rescue experiments.
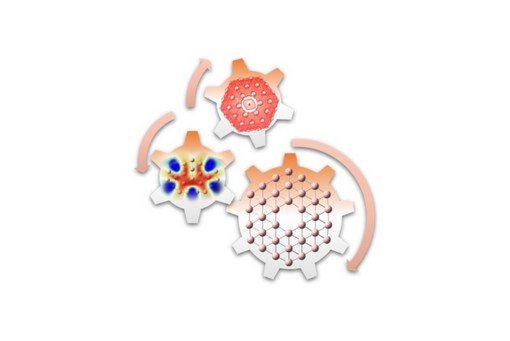
In continuation of our previous work of small boron clusters [Phys. Chem. Chem. Phys. 23, 24118 (2021)], we systematically examined the neutral and negatively charged boron clusters Bn/B'n − (nε{8–38}) with planar or quasi-planar conformations, whose exotic properties have aroused considerable interest in the literature.
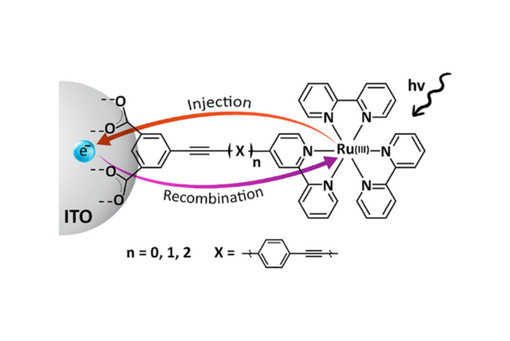
Here, LL is a 4-substituted 2,2-bipyridine (bpy) ligand with varying numbers of conjugated phenylenethynylene bridge units between the bipyridine ring and anchoring group consisting of a bis-carboxylated isophthalic group.
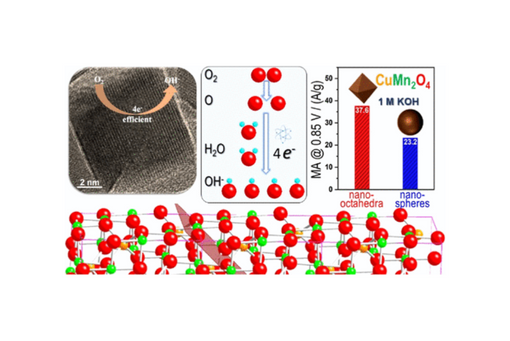
To further exploit the potential of non-Pt-group metal-based spinel catalysts for the alkaline oxygen reduction reaction (ORR), a cathodic fuel cell reaction, we hereby report a strategy of ORR improvement by controlling the crystallographic facets of ultra-small CuMn2O4 spinel nanocatalysts through a developed colloidal synthesis approach.
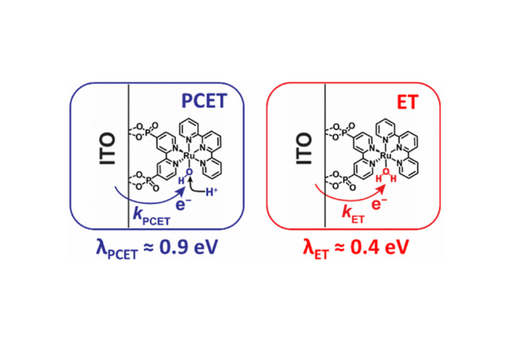
The reorganization energy (λ) for interfacial electron transfer (ET) and proton-coupled ET (PCET) from a conductive metal oxide (In2O3:Sn, ITO) to a surface-bound water oxidation catalyst was extracted from kinetic data measured as a function of the thermodynamic driving force.
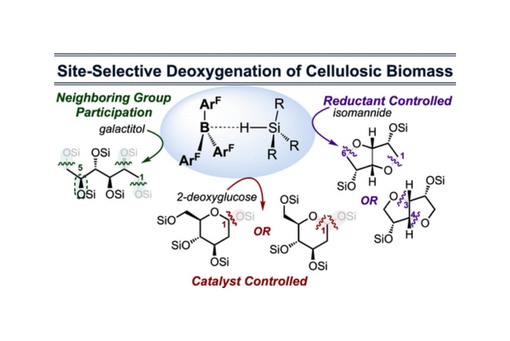
This perspective describes key lessons from a program whose goals are to site-selectively deoxygenate cellulose-derived carbohydrates.

We measured airborne PFAS on PM2.5 filters in close proximity to a major fluoropolymer manufacturing facility (Chemours' Fayetteville Works) located near Fayetteville, North Carolina, USA.
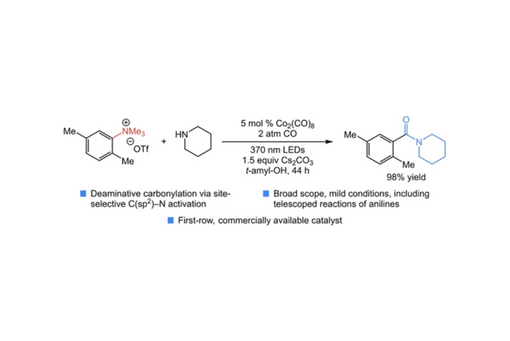
Herein, we demonstrate an aminocarbonylation of aniline-derived trialkylammonium salts promoted by visible light with a simple cobalt catalyst.
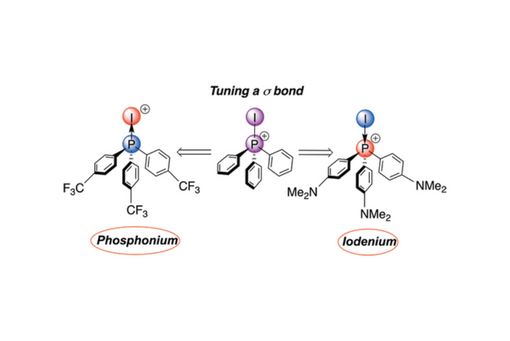
Herein, we use an ensemble of computational tools and methodologies to probe the nature of this ambi-valent bond.
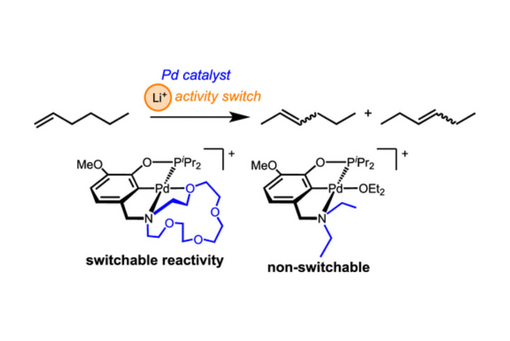
Palladium(II) pincer complexes with varying amine donor substituents have been prepared and studied in olefin isomerization catalysis.
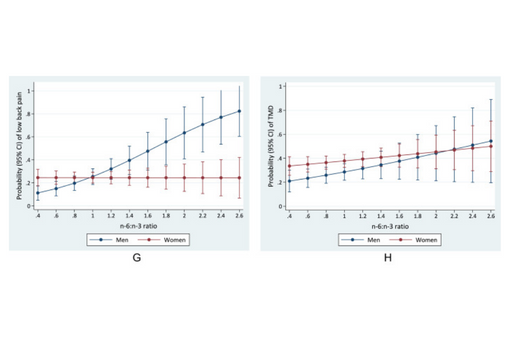
This cross-sectional study investigated associations between the omega-6/omega-3 PUFA ratio and somatic symptom disturbance and depressive symptoms in a community-based sample of 501 adults and determined whether these associations differed between adults with and without TMD or irritable bowel syndrome (IBS).