Structure and Function of a Dehydrating Condensation Domain in Nonribosomal Peptide Biosynthesis
Abstract
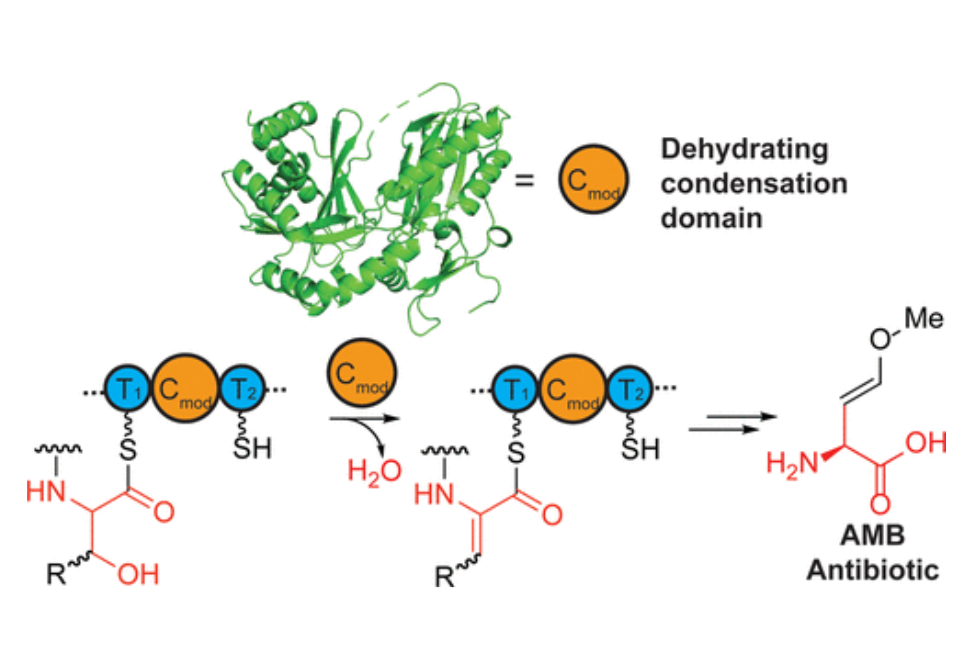
Dehydroamino acids are important structural motifs and biosynthetic intermediates for natural products. Many bioactive natural products of nonribosomal origin contain dehydroamino acids; however, the biosynthesis of dehydroamino acids in most nonribosomal peptides is not well understood. Here, we provide biochemical and bioinformatic evidence in support of the role of a unique class of condensation domains in dehydration (CmodAA). We also obtain the crystal structure of a CmodAA domain, which is part of the nonribosomal peptide synthetase AmbE in the biosynthesis of the antibiotic methoxyvinylglycine. Biochemical analysis reveals that AmbE-CmodAA modifies a peptide substrate that is attached to the donor carrier protein. Mutational studies of AmbE-CmodAA identify several key residues for activity, including four residues that are mostly conserved in the CmodAA subfamily. Alanine mutation of these conserved residues either significantly increases or decreases AmbE activity. AmbE exhibits a dimeric conformation, which is uncommon and could enable transfer of an intermediate between different protomers. Our discovery highlights a central dehydrating function for CmodAA domains that unifies dehydroamino acid biosynthesis in diverse nonribosomal peptide pathways. Our work also begins to shed light on the mechanism of CmodAA domains. Understanding CmodAA domain function may facilitate identification of new natural products that contain dehydroamino acids and enable engineering of dehydroamino acids into nonribosomal peptides.
Citation
Structure and Function of a Dehydrating Condensation Domain in Nonribosomal Peptide Biosynthesis
DOI: 10.1021/jacs.1c13404