Spatially resolved quantification of drug metabolism and efficacy in 3D paper-based tumor mimics
Abstract
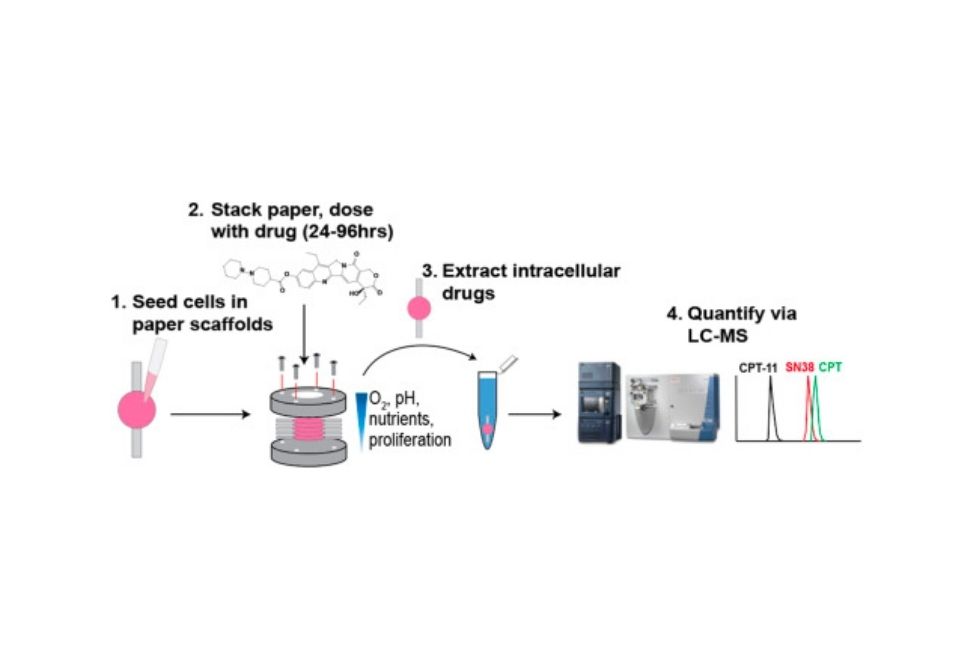
Paper-based cultures are an emerging platform for preparing three-dimensional (3D) tissue- and tumor-like structures. The ability to stack individual sheets of cell-containing paper affords a modular means of assembling structures with defined cellular compositions and microenvironments. These layered stacks are easily separated at the end of an experiment, providing spatially resolved populations of live cells for further analysis. Here we describe a workflow in which cell viability, drug penetration, and drug metabolism are quantified in a spatially resolved manner. Specifically, we mapped the distribution of the drug irinotecan and its bioactive metabolite SN38 in a colorectal cancer cell-containing stacked structure with liquid chromatography-mass spectrometry (LC-MS). This paper provides the first example of a 3D culture platform that quantifies viability and drug metabolism in a spatially resolved manner. Our data show that cells at the bottom of the stack are more drug-resistant than layers in contact with the culture medium, similar to cells in the nutrient-poor center of a proliferating tumor being more drug-resistant than the rapidly dividing cells at its periphery. The powerful combination of quantitative viability and drug metabolism measurements will enable future studies to determine the exact mechanism(s) of drug resistance in different regions of a tumor.
Citation
Tyler S. Larson, Gary L. Glish, Matthew R. Lockett,
Spatially resolved quantification of drug metabolism and efficacy in 3D paper-based tumor mimics,
Analytica Chimica Acta,
Volume 1186,
2021,
339091,
ISSN 0003-2670,
https://doi.org/10.1016/j.aca.2021.339091.