Contributions of methionine to recognition of trimethyllysine in aromatic cage of PHD domains: implications of polarizability, hydrophobicity, and charge on binding
Abstract
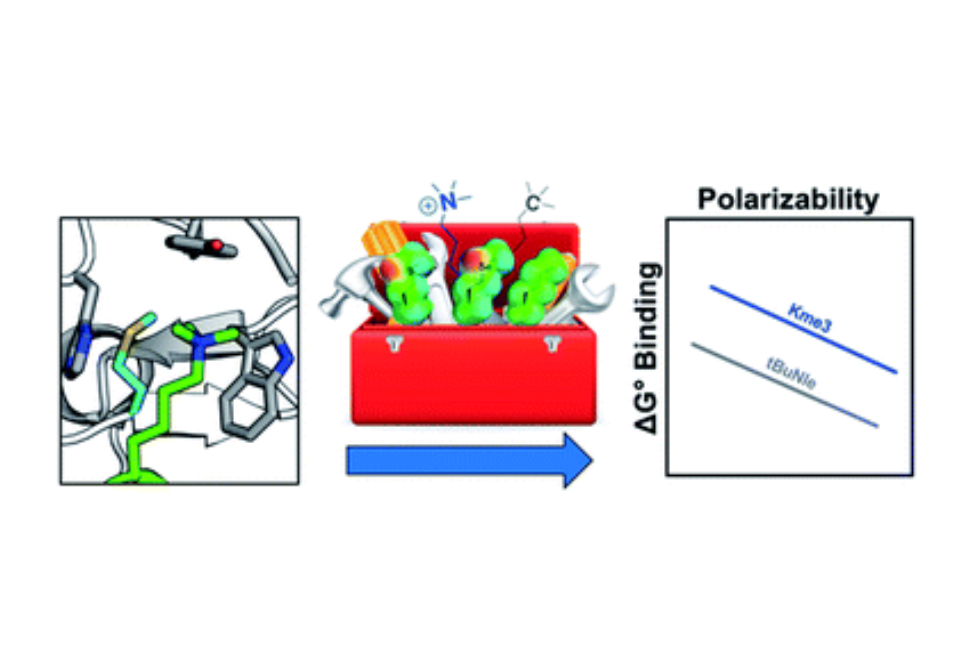
Recognition of trimethyllysine (Kme3) by reader proteins is an important regulator of gene expression. This recognition event is mediated by an aromatic cage made up of 2–4 aromatic residues in the reader proteins that bind Kme3 via cation–π interactions. A small subset of reader proteins contain a methionine (Met) residue in place of an aromatic sidechain in the binding pocket. The unique role of sulfur in molecular recognition has been demonstrated in a number of noncovalent interactions recently, including interactions of thiols, thioethers, and sulfoxides with aromatic rings. However, the interaction of a thioether with an ammonium ion has not previously been investigated and the role of Met in binding Kme3 has not yet been explored. Herein, we systematically vary the Met in two reader proteins, DIDO1 and TAF3, and the ligand, Kme3 or its neutral analog tert-butyl norleucine (tBuNle), to determine the role of Met in the recognition of the cationic Kme3. Our studies demonstrate that Met contributes to binding via dispersion forces, with about an equal contribution to binding Kme3 and tBuNle, indicating that electrostatic interactions do not play a role. During the course of these studies, we also discovered that DIDO1 exhibits equivalent binding to tBuNle and Kme3 through a change in the mechanism of binding.
Citation
Contributions of methionine to recognition of trimethyllysine in aromatic cage of PHD domains: implications of polarizability, hydrophobicity, and charge on binding
Katherine I. Albanese and Marcey L. Waters
Chemical Science 2021, Advance Article
DOI: 10.1039/d1sc02175c