A structural metagenomics pipeline for examining the gut microbiome
Abstract
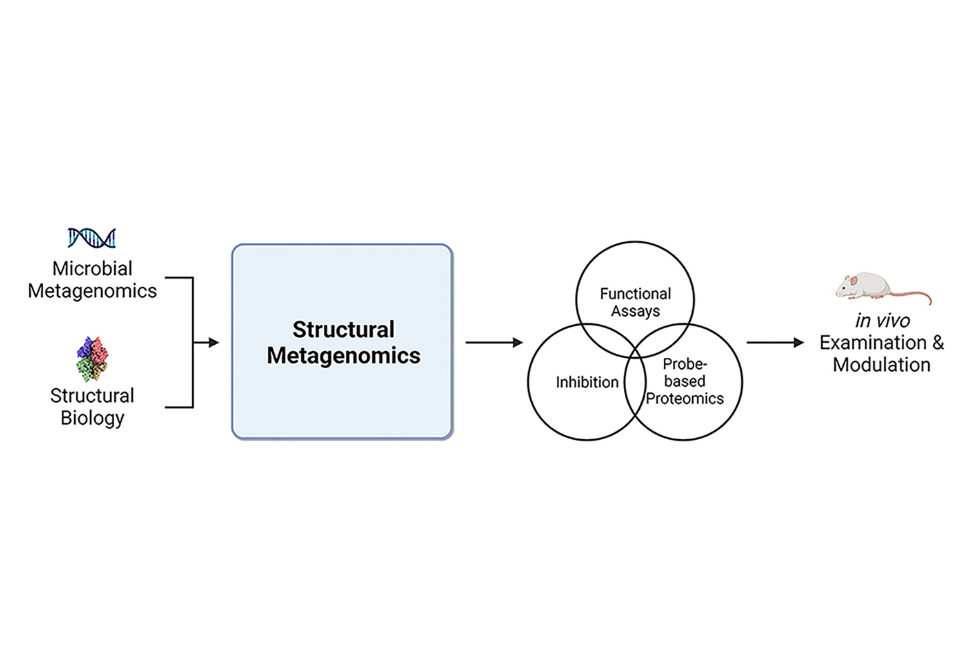
Metagenomic sequencing data provide a rich resource from which to expand our understanding of differential protein functions involved in human health. Here, we outline a pipeline that combines microbial whole genome sequencing with protein structure data to yield a structural metagenomics-informed atlas of microbial enzyme families of interest. Visualizing metagenomics data through a structural lens facilitates downstream studies including targeted inhibition and probe-based proteomics to define at the molecular level how different enzyme orthologs impact in vivo function. Application of this pipeline to gut microbial enzymes like glucuronidases, TMA lyases, and bile salt hydrolases is expected to pinpoint their involvement in health and disease and may aid in the development of therapeutics that target specific enzymes within the microbiome.
Citation
Morgan E. Walker, Joshua B. Simpson, Matthew R. Redinbo,
A structural metagenomics pipeline for examining the gut microbiome,
Current Opinion in Structural Biology,
Volume 75, 2022, 102416, ISSN 0959-440X,
https://doi.org/10.1016/j.sbi.2022.102416.
(https://www.sciencedirect.com/science/article/pii/S0959440X22000951)