Single-Molecule Correlated Chemical Probing: A Revolution in RNA Structure Analysis
Abstract
RNA molecules convey biological information both in their linear sequence and in their base-paired secondary and tertiary structures. Chemical probing experiments, which involve treating an RNA with a reagent that modifies conformationally dynamic nucleotides, have broadly enabled examination of short- and long-range RNA structure in diverse contexts, including in living cells. For decades, chemical probing experiments have been interpreted in a per-nucleotide way, such that the reactivity measured at each nucleotide reports the average structure at a position over all RNA molecules within a sample. However, there are numerous important cases where per-nucleotide chemical probing falls short, including for RNAs that are bound by proteins, RNAs that form complex higher order structures, and RNAs that sample multiple conformations.
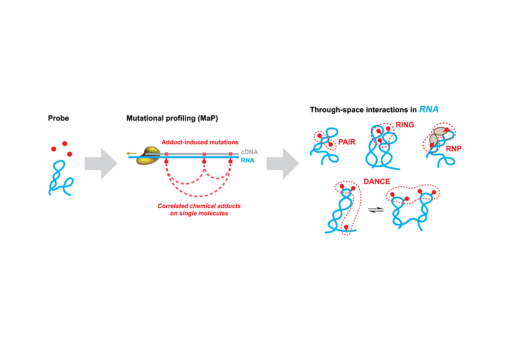
Recent experimental and computational innovations have started a revolution in RNA structure analysis by transforming chemical probing into a massively parallel, single-molecule experiment. Enabled by a specialized reverse transcription strategy called mutational profiling (MaP), multiple chemical modification events can be measured within individual RNA molecules. Nucleotides that communicate structurally through direct base pairing or large-scale folding–unfolding transitions will react with chemical probes in a correlated manner, thereby revealing structural complexity hidden to conventional approaches. These single-molecule correlated chemical probing (smCCP) experiments can be interpreted to directly identify nucleotides that base pair (the PAIR-MaP strategy) and to reveal long-range, through-space structural communication (RING-MaP). Correlated probing can also define the thermodynamic populations of complex RNA ensembles (DANCE-MaP). Complex RNA–protein networks can be interrogated by cross-linking proteins to RNA and measuring correlations between cross-linked positions (RNP-MaP).
smCCP thus visualizes RNA secondary and higher-order structure with unprecedented accuracy, defining novel structures, RNA–protein interaction networks, time-resolved dynamics, and allosteric structural switches. These strategies are not mutually exclusive; in favorable cases, multiple levels of RNA structure ─ base pairing, through-space structural communication, and equilibrium ensembles ─ can be resolved concurrently. The physical experimentation required for smCCP is profoundly simple, and experiments are readily performed in cells on RNAs of any size, including large noncoding RNAs and mRNAs. Single-molecule correlated chemical probing is paving the way for a new generation of biophysical studies on RNA in living systems.
Citation
Single-Molecule Correlated Chemical Probing: A Revolution in RNA Structure Analysis Anthony M. Mustoe, Chase A. Weidmann, and Kevin M. Weeks Accounts of Chemical Research 2023 56 (7), 763-775 DOI: 10.1021/acs.accounts.2c00782