Marcey Waters
Glen H. Elder, Jr. Distinguished Professor
Caudill Laboratories 219919-843-6522
mlwaters@email.unc.edu
Group Website
Curriculum Vitae
Research Interests
Bioorganic Chemistry, Molecular Recognition
Research Synopsis
Our group is an interdisciplinary group, focusing on problems of molecular and biomolecular recognition. Molecular recognition impacts a wide range of fields, including asymmetric catalysis, materials chemistry, and protein folding. Consider, for example, designing a drug to bind to the active site of an enzyme. What features other than shape might contribute to binding? What types of interactions will provide high affinity as well as high selectivity? These are general questions in the field of molecular recognition that we are investigating for applications to biosensing, drug delivery, and de novo protein design. The research interests in our group span a wide range, from mechanistic organic chemistry and molecular recognition to structural biology, and hence involve the use of a variety of techniques. Methods used in our group include organic and solid phase synthesis, combinatorial chemistry, computational chemistry, molecular biology, kinetic and thermodynamic measurements using 1D and 2D NMR, circular dichroism, UV/Vis and fluorescence spectroscopy, analytical ultracentrifugation, and calorimetry. The extent that any one student uses these techniques depends largely on the particular student's research interests.
Professional Background
BA, UCSD, 1992; PhD, The University of Chicago, 1997; NIH Postdoctoral Fellow, Columbia University, 1997-1999; NSF Career Award, 2001-2006; Alfred P. Sloan Fellow, 2004-2006; Board of Directors, Mesilla Chemistry Workshop; Advisory Board Member, International Symposia on Macrocyclic and Supramolecular Chemistry.
Research Group
News & Publications

Chemistry Ph.D. graduate Lindsey James is developing chemical tools to study how cells decide which genes to use, when to use them and when to keep them silent.
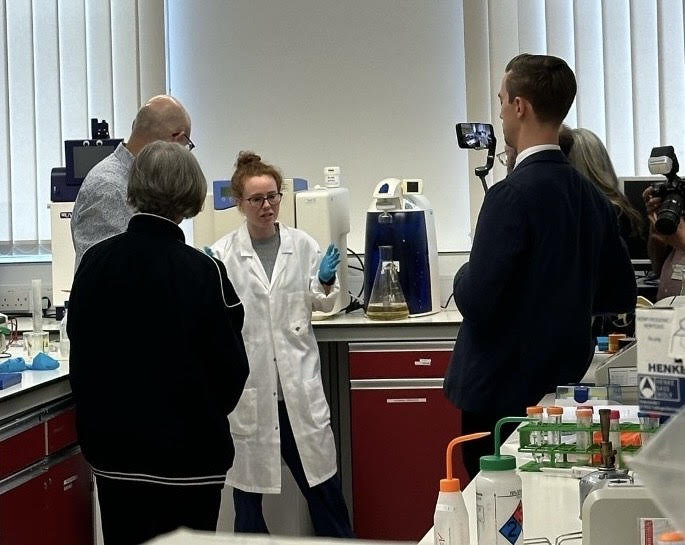

In her lab at Wake Forest University, Assistant Professor Katherine Albanese is reimagining proteins as tools for probing life at the molecular level—decoding how subtle chemical tags on DNA-packaging proteins affect gene expression and, ultimately, health.